Spatial transcriptomics reveals ovarian cancer subclones with distinct tumour microenvironments
- To whom correspondence should be addressed: Elena Denisenko, Yu Yu, Paul Cohen, or Alistair Forrest
- Harry Perkins Institute of Medical Research, QEII Medical Centre and Centre for Medical Research, The University of Western Australia, Nedlands, Perth, WA 6009, Australia
- Children’s Hospital of Eastern Ontario Research Institute, Ontario, Canada (Current Affiliation)
- Anatomical Pathology Department, Clinipath, Sonic Healthcare, Perth, WA 6017 Australia
- Centre for Precision Health, Edith Cowan University, Joondalup, WA 6027, Australia
- Department of Gynaecological Oncology, Bendat Family Comprehensive Cancer Centre, St John of God Subiaco Hospital, 12 Salvado Rd, Subiaco, WA 6008, Australia
- School of Medicine, University of Notre Dame, Fremantle,WA,6160, Australia
- Western Australian Gynae and Surgery, Perth, WA, Australia.
- Division of Obstetrics and Gynaecology, Medical School, University of Western Australia, 35 Stirling Highway, Crawley, WA 6009, Australia
- Institute for Health Research, The University of Notre Dame Australia, 32 Mouat Street Fremantle, WA 6160, Australia
- Curtin Medical School, Curtin University, 410 Koorliny Way, Bentley, WA 6102, Australia
- Curtin Health Innovation Research Institute, Curtin University B305, Bentley, WA 6102, Australia
Abstract
High-grade serous ovarian carcinoma (HGSOC) is characterised by recurrence, chemotherapy resistance and overall poor prognosis. Genetic heterogeneity of tumour cells and the microenvironment of the tumour have been hypothesised as key determinants of treatment resistance and relapse. Here, using a combination of spatial and single cell transcriptomics (10x Visium and Chromium platforms), we examine tumour genetic heterogeneity and infiltrating populations of HGSOC samples from eight patients with variable response to neoadjuvant chemotherapy. By inferring gross copy number alterations (CNAs), we identified distinct tumour subclones co-existing within individual tumour sections. These tumour subclones have unique CNA profiles and spatial locations within each tumour section, which were further validated by ultra-low-pass whole genome sequencing. Differential expression analysis between subclones within the same section identified both tumour cell intrinsic expression differences and markers indicative of different infiltrating cell populations. The gene sets differentially expressed between subclones were significantly enriched for genes encoding plasma membrane and secreted proteins, indicative of subclone-specific microenvironments. Furthermore, we identified tumour derived ligands with variable expression levels between subclones that correlated or anticorrelated with various non-malignant cell infiltration patterns. We highlight several of these that are potentially direct tumour-stroma/immune cell relationships as the non-malignant cell type expresses a cognate receptor for the tumour derived ligand. These include predictions of CXCL10-CXCR3 mediated recruitment of T and B cells to associate with the subclones of one patient and CD47-SIRPA mediated exclusion of macrophages from association with subclones of another. Finally, we show that published HGSOC molecular subtype signatures associated with prognosis are heterogeneously expressed across tumour sections and that areas containing different tumour subclones with different infiltration patterns can match different subtypes. Our study highlights the high degree of intratumoural subclonal and infiltrative heterogeneity in HGSOC which will be critical to better understand resistance and relapse.
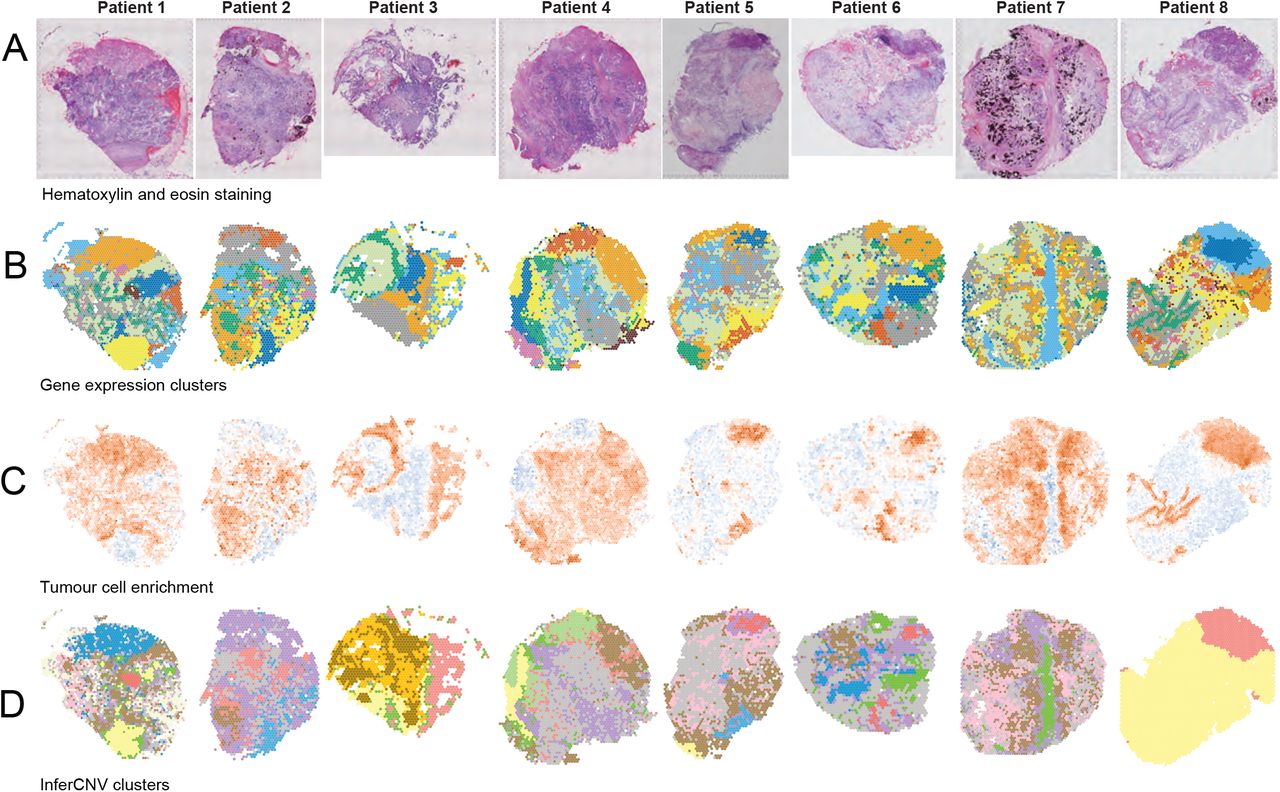
Visium profiles of eight HGSOC samples.
Shown are a) hematoxylin and eosin stained tissue sections, b) gene expression-based clusters mapped onto the tissue sections, c) tumour cell enrichment scores calculated using Giotto 29. Red indicates enrichment, blue depletion, d) InferCNV clusters. Yellow spots in patients 1, 3, 4, 5, 8 correspond to non-malignant regions, green corresponds to border regions, other colours correspond to putative tumour subclones.